Cellular metabolism
Cellular metabolism with its more than 1000 different chemical reactions is highly complex. Despite decades of research, we are still far from understanding how metabolism really functions. Yet, such understanding is crucial for biotechnology as well as for health and disease, where we start to realize that a faulty metabolism might be at the onset of diseases such as diabetes and cancer, and of microbial pathogenicity etc. Also, in recent years, we have more and more evidence that metabolism is not a boring cellular function, but that metabolism is in fact also involved in cellular decision-making.
Our aims
To this end, our lab wants to understand (i) how primary carbon metabolism functions, and (ii) how metabolism controls other cellular processes, such as the entry into bacterial persistence or progression through the eukaryotic cell cycle.
Our approach
In our projects, we combine experimental and computational/modeling efforts. Where necessary, we develop new experimental or computational tools. Our lab members have diverse educational backgrounds ranging from biology via engineering to computer science. We find such a multidisciplinary environment very stimulating and also most adequate to jointly solve the exciting challenges.
Our research is primarily curiosity-driven, i.e. the main source of motivation comes from the mere interest of getting a deeper insight into metabolism. However, in case one of our generated fundamental findings can, for instance, help improving bioprocesses, we also try to implement this in the applied context.
How does primary metabolism function?
We have discovered two important principles of how metabolism works:
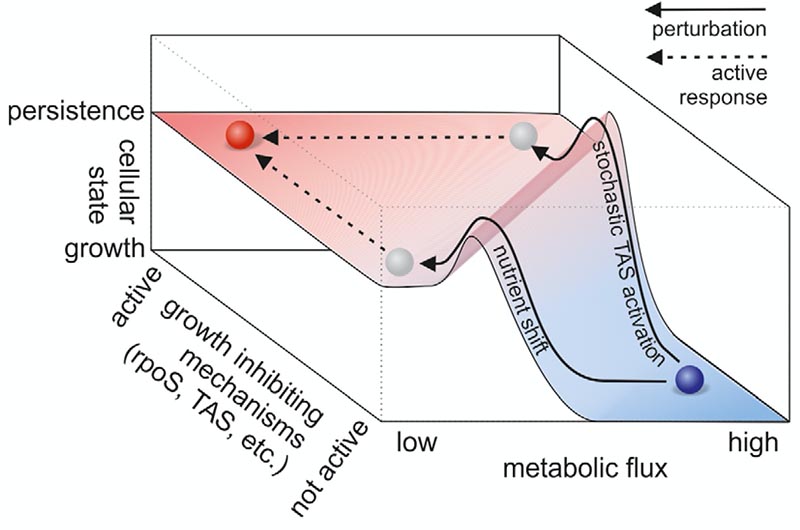
First, we uncovered – initially by computational predictions (Kotte et al, Mol Sys Biol, 2010) and later validated by experiments (Kochanowski et al, PNAS, 2013) – that cells have mechanisms in place that sense metabolic flux and that are used for flux-dependent regulation. Further, we found that flux-sensing and flux-dependent regulation can lead to responsive diversification of clonal E. coli populations into multiple phenotypes, where one of these can be the medically relevant persister phenotype (Kotte et al, Mol Sys Biol, 2014; Radzikowski et al, Mol Sys Biol, 2016), and also can have relevance during yeast aging (Hubmann et al, eLife, 2019).
Second, recently, we found that cells are limited by the rate with which they dissipate Gibbs energy (Niebel et al, Nature Metabol, 2019). We found this by developing a constraint-based model for Saccharomyces cerevisiae with a comprehensive description of biochemical thermodynamics, and by using non-linear regression analyses of quantitative metabolome and physiology data. This finding opens up a completely new avenue on how metabolism eventually controls cellular functioning.
How does metabolism control other cellular functions?
In recent years, we worked on two examples where metabolism controls other cellular processes: the eukaryotic cell cycle and the entry into persistence in bacteria:
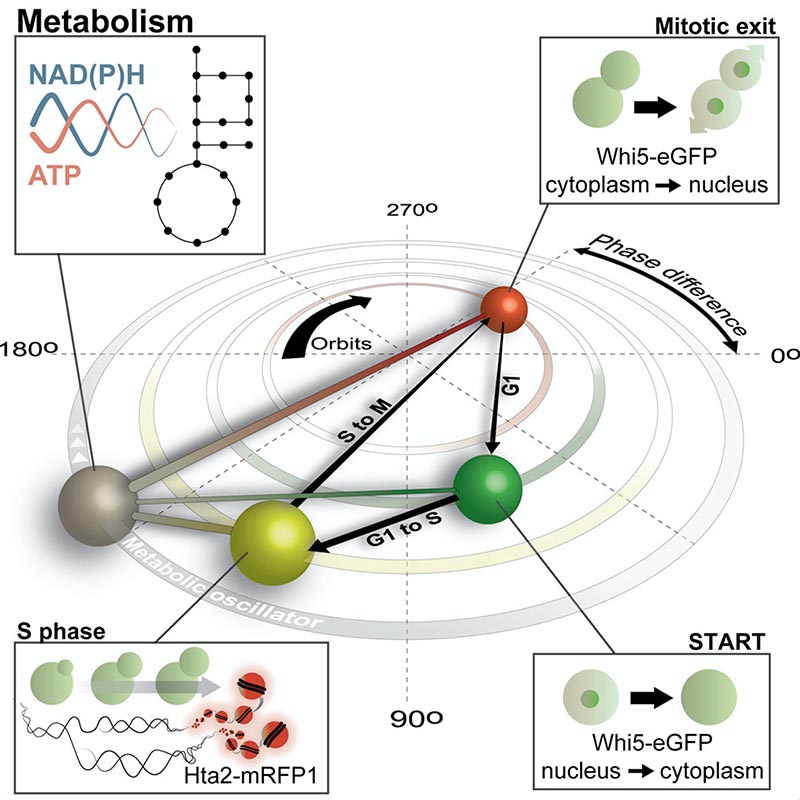
Using dynamic single-cell metabolite measurements, we found that metabolism of yeast is an autonomous oscillator (Papagiannakis et al, Mol Cell, 2017). Offering a new view on how cell cycle control might be exerted, through solving a complex inverse problem, we recently obtained evidence that the metabolic oscillator could in fact control the early and late cycle independently (Özsezen et al, Cell Syst, 2019). Through highly sophisticated dynamic microscopic analyses, we found that protein synthesis rates increase during the G1 phase, causing the levels of the early cyclin Cln3 to increase and leading to the cell cycle start (Litsios et al, Nature Cell Biol, 2019).
In E. coli, we found that upon abrupt nutrient shifts, a homogeneous population can diversify into two populations (Kotte et al, Mol Syst Biol, 2014). One of the populations that emerges is composed of persister cells, i.e. cells that are tolerant to antibiotics without carrying any genetically-determined resistance. We found that this persister state still has an active metabolism and is geared towards a stress response (Radzikowski et al, Mol Syst Biol, 2016).
